Jupyter Notebooks
Jupyter Notebooks emerged from the IPython Notebook [Perez2007]. A notebook document consists of a series of "cells", which can either contain computer code or markdown text. For those with computer code, the code in the cell can be executed (by being sent to a so-called "kernel", evaluated, and results and output returned to the notebook), and the output from that execution is shown just below in the same cell.
Here is a typical example with a Python kernel:
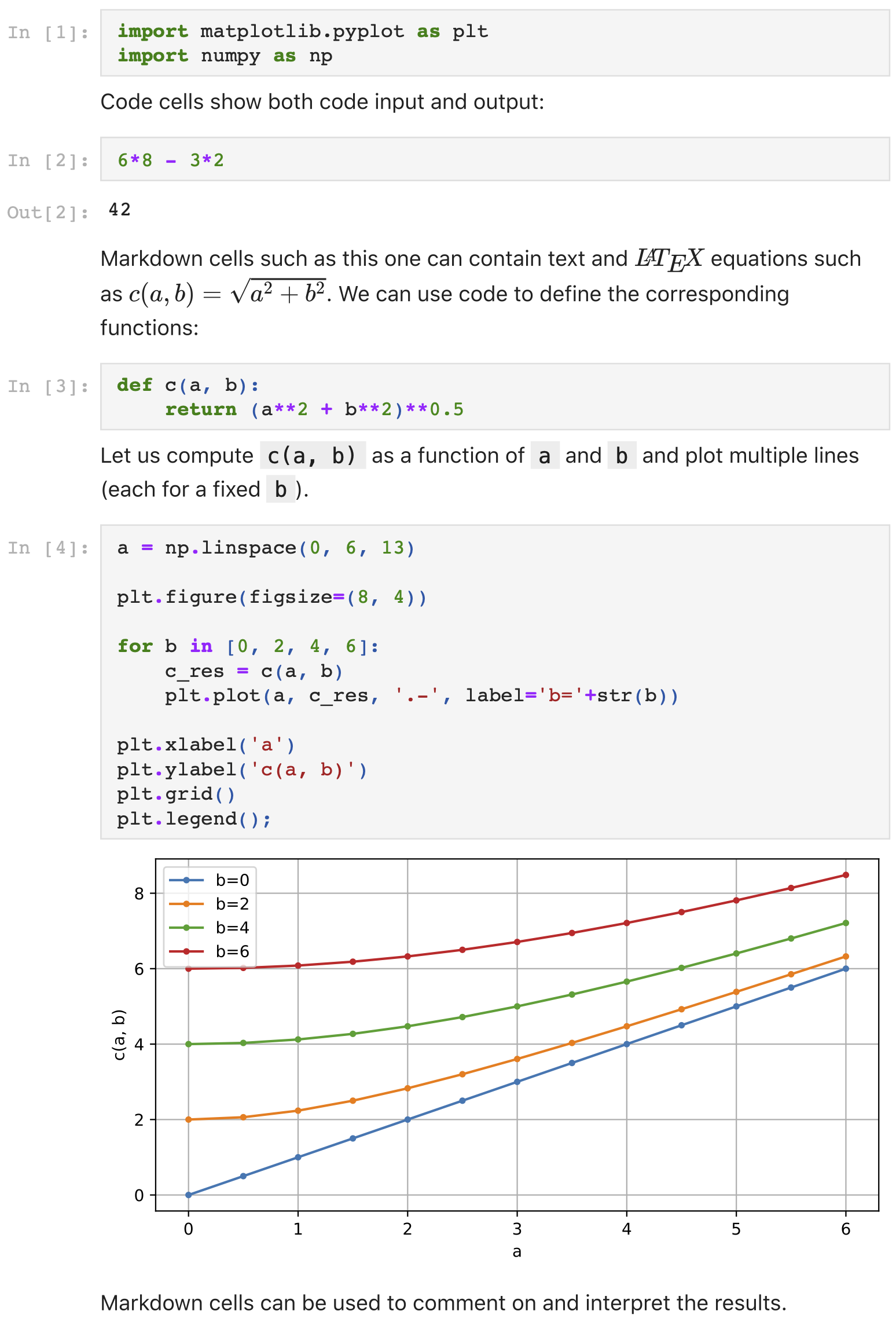
For more details, see the home page of project Jupyter.
Or try the notebook in your browser using https://jupyter.org/try .
Use cases in Science
Notebooks help to explore a data set interactively. Each executed code cell is an attempt by the researcher to achieve something and to tease out some insight from the data set. The result is displayed immediately below the code commands, and the researcher can pause and think about the outcome. As code cells can be executed in any order, modified and re-executed as desired, deleted and copied, the notebook is a convenient environment to iteratively explore a complex problem.
Over time the researchers keep the cells that contain good steps (with added text cells for explanation and discussion) while discarding all attempts that did not lead to anything.
Brian Granger and Fernando Perez, two of the co-leaders of Project Jupyter, describe this as "Jupyter helps humans to think and tell stories with code and data" [Granger2021].
At the end of a data exploration session (or the end of the day of work), the notebook can be saved, and when the researcher is ready to continue, they load the notebook document again, execute all cells stored in it so far, and continue the investigation.
The above example refers to the exploration of a data set, but the same use of notebooks is of value when carrying out simulation based studies, where the simulation may be driven from the notebook, and the simulation results analysed from the same notebook [Fangohr2020, Beg2021]. Similarly, computational exploration of mathematics can be done from the notebook [Beg2021].
Notebooks can tell the story of a scientific exploration for others: narrative (in markdown format, including LaTeX if desired) can be combined with code that processes data and creates plots from it which are shown in the notebook. Additional text can be used to interpret the data. These are the essentials of a scientific manuscript, combined in one document. This is described as "one-study one-document" in [Beg2021].
Notebooks can be shared easily and simplify communication with co-workers and supervisors: Notebooks can be converted to html and pdf, and then shared (by email for example) as static read-only documents. This is useful to communicate and share a study with colleagues or managers. By adding sufficient explanation, the main story can be understood by the reader, even if they wouldn't be able to write the code that is embedded in the document. Nevertheless, the description is complete as the code must contain all the information required to compute the outputs shown in the notebook. (Where code cells become very large, it is good practice to gather relevant functionality and store it in an external file - in the case of Python in a module.py file or a package which can then be imported from the notebook [Fangohr2020].)
Notebooks can be used interactively or as a script: a common pattern in science is that a computational recipe is iteratively developed in a notebook. Once this has been found and should be applied to further data sets (or other points in some parameter space), the notebook can be executed like a script, for example by submitting these scripts as batch jobs to a High Performance Computing installation. This is described in [Beg2021], and [Fangohr2020] presents this "notebook-as-a-script" approach in the context of the European X-ray Free Electron Laser research facility.
A collection of notebook recipes can be used as a library for typical computational tasks in a particular scientific domain [Fangohr2020]. For use of such computational recipes in a research context, there needs to be the flexibility to easily change the recipe: it is core to research that new things are studied, and thus existing recipes may be a good starting point or a nearly perfect solution but are likely to need small changes. A recipe stored in a Jupyter notebook makes this somewhat convenient, as a copy of a recipe can be used as a starting point, and then some of the cells can be modified as is required to adapt the recipe to the new computational task at hand.
Remote use of High Performance Compute resources: the computational kernel used by a notebook can be hosted on the same machine as the browser that displays the notebook client, but does not have to be. Using JupyterHub, notebooks can be executed on the computing resources of a research facility, supercomputing centre or university, while the researcher driving it does this from a web browser on their laptop [Fangohr2020]. The access to HPC resources via Notebooks and JupyterHub is often a sufficient replacement of remote X sessions. Remote access to is also useful if the amount of data to process is significant, and the data cannot be moved easily [Fangohr2020].
The Binder project [Jupyter2018] exposes remote computing resources like JupyterHub, but creates the software environment in which the execution takes place on demand (using repo2docker) and specifically for each notebook. This makes it possible to store a set of notebooks together with a specification of their required software environment in a public repository on github such as this demonstration repository. The public mybinder.org service can then be used to interactively execute and control these notebooks in the software environment they require, using cloud resources provided (indirectly) by mybinder.org.
Use cases for mybinder include provision of (interactive) documentation of software that can be executed if desired, services which allow to try a software or analysis in the cloud (such as Jupyter's own https://jupyter.org/try service), and (interactive) textbooks, where each chapter is given through a notebook and can be interactively executed and studied by the student - for example this textbook on Python programming. Another use case is the provision of reproducible procedures:
Notebooks can help to achieve reproducible scientific results. If a notebook is used to drive the data analysis - starting from raw data or simulation results - to the creation of publication ready figures, then by preserving the notebook, we have documented the computational procedure. Following this (slightly simplified) idea, one can make publications more reproducible, by accompanying publications with a set of notebooks that, when executed, reproduce essential components of the paper, such as figures, tables or important numbers [Beg2021]. An example for such additional material to a paper by Max Albert is this git repository: the notebooks subdirectory contains notebooks that create all the figures in this publication by Maximilian Albert or the LIGO experiment [Kluyver2016].
It has been proposed that some of the Vision of the European Open Science Cloud could be realised through Jupyter Notebooks, JupyterHub and BinderHubs [Fangohr2020, Fangohr2022].
Why such a success?
The recent paper "Jupyter: Thinking and Storytelling with Code and Data" by Brian Granger and Fernando Perez [Branger2021] contains a number of interesting perspectives and insights why the Jupyter notebook has quickly gained traction in academia and (maybe even more so) elsewhere. One of the thoughts is that the design of the notebook focuses on the human: "Jupyter helps individuals and groups to leverage computation and data to solve complex, technical, but human-centered problems of understanding, decision making, collaboration, and community practice".
Feedback
The above selection of use cases and references to publications is incomplete and subjective. If you are aware of additional related publications or use cases, please get in touch.
References
[Perez2007] Fernando Perez and Brian E. Granger, "IPython: A System for Interactive Scientific Computing" in Computing in Science & Engineering, vol. 9, no. 3, pp. 21-29, May-June 2007, doi: 10.1109/MCSE.2007.53. http://fperez.org/papers/ipython07_pe-gr_cise.pdf (2007)
[Kluyver2016] Thomas Kluyver, Benjamin Ragan-Kelley, Fernando Pérez, Brian Granger, Matthias Bussonnier, Jonathan Frederic, Kyle Kelley, et al, "Jupyter Notebooks – a publishing format for reproducible computational workflows", https://doi.org/10.3233/978-1-61499-649-1-87, http://ebooks.iospress.nl/publication/42900 (2016)
[Jupyter2018] Jupyter et al., "Binder 2.0 - Reproducible, Interactive, Sharable Environments for Science at Scale." Proceedings of the 17th Python in Science Conference. 2018. http://doi.org/10.25080/Majora-4af1f417-011 (2018)
[Fangohr2020] Hans Fangohr, Marijan Beg, Martin Bergemann, Valerii Bondar, Sandor Brockhauser, Camille Carinan, Raul Costa, et al, "Data exploration and analysis with Jupyter notebooks", Proceedings of the 17th International Conference on Accelerator and Large Experimental Physics Control Systems ICALEPCS2019, TUCPR02 http://accelconf.web.cern.ch/icalepcs2019/doi/JACoW-ICALEPCS2019-TUCPR02.html, pdf (2020)
[Beg2021] Marijan Beg; Juliette Belin; Thomas Kluyver; Alexander Konovalov; Min Ragan-Kelley; Nicolas Thiery; Hans Fangohr. "Using Jupyter for Reproducible Scientific Workflows", Computing in Science & Engineering, vol. 23, no. 2, pp. 36-46, 1 March-April 2021, doi: 10.1109/MCSE.2021.3052101 arXiv preprint (2021)
[Granger2021] Brian Granger, Fernando Pérez. "Jupyter: Thinking and Storytelling With Code and Data." Computing in Science & Engineering, vol. 23, no. 2, pp. 7-14, 1 March-April 2021, doi: 10.1109/MCSE.2021.3059263 Authorea preprint (2021)
[Fangohr2022] Hans Fangohr. Presentation "EOSC Jupyter Vision", 29 November 2022, Grenoble, France (pdf) (2022)
History
- Original post from 9 April 2021.
- Updated 30 November 2022: - add last reference - add URLs in main text (and DOIs in some places)